Research into protein folding can lead to medical breakthroughs
A Wilkes University research team headed by Dr. Del Lucent, associate professor of physics and Dr. Sofya Chepushtanova, assistant professor of mathematics, is gaining attention for its study on protein folding.
The team hopes to build models from computer software that can predict how a protein folds and where it can go wrong.
Protein folding is a fundamental piece of how every living thing works. It occurs when a protein is created and folding makes it contort into its final shape.
All the things that happen in your cells, with a small number of exceptions, are mediated by proteins doing their jobs. They are the building blocks of biology.
Most people think of proteins as the macronutrient, especially with regard to lifting weights.
While this is correct, the rest of you is also made of protein.
There are many different kinds of proteins. Some move objects from one location to the other. Others transfer signals from one location in a cell to the other. Others catalyze chemical reactions.
When a protein folds incorrectly, it can cause diseases such as Alzheimer’s disease, Huntington’s disease or Parkinson’s disease. Many genetic diseases are the result of mistakes in the instructions for protein folding. For example, a mistake in the protein that’s responsible for hemoglobin assembly results in sickle cell anemia.
“Let’s say a protein would take a millisecond to fold. That means in order to reach that step, we need to break that up into steps of a femtosecond in size. We would need to break that up into basically a thousand billion steps. So it’s like measuring the distance from here to the Moon one millimeter at a time,” Lucent said.
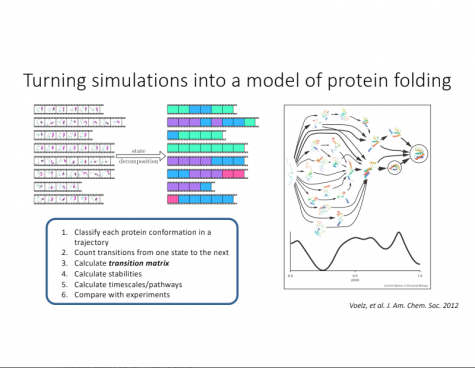
Proteins are too small, and their folding occurs too fast, to be observed naturally. Therefore, it is necessary to create computer simulations to observe these changes.
To get an accurate model of how a protein moves, all of the pieces of a protein need to be moved in very small amounts many times. This results in data sets and simulations that are multiple terabytes in length.
“All these calculations are not very fast. So if you compute this barcode, take persistence imaging, then you apply a machine-learning algorithm to that, it can take years. Some of the students even made calculations for that,” Chepushtanova said.
Joseph Gubbiotti, senior computer science major, was one of those students. He wrote a computer program to calculate the barcodes for non-hydrogen atoms. For a (relatively) small protein, ACBP, the calculations would take upwards of 20 years to complete.
“The goal of this research group is to determine if focusing on particularly interesting mathematical features of proteins would produce a computationally less expensive way of predicting protein folding. Simulating proteins folding using molecular dynamics is possible, but incredibly time and resource consuming,” Gubbiotti said.
Topological data analysis is used to distill these massive data sets and compartmentalize them into groups based on their shapes.
The study of topology is observing the properties of shapes, specifically shape deformation without tearing or gluing. These shapes can be continuously bent and stretched so long as they retain the same number of holes. It is useful in studying the motion of a protein in the process of folding.
“We’re very close to submitting something for publication with regards to topological data analysis as a proof of concept. If we can figure out how misfolding happens and where we can intercede, we can come up with a way to rescue that misfolded protein,” Lucent said. “That’s the goal of the research, and I hope to have broken ground and made some progress towards that goal over the next year.”

The six monitor setup in Lucent’s lab requires multiple high-end GPUs to power the monitors and the computer simulations. It has 126 gigabytes of RAM, over two terabytes of storage and two 8-core CPUs. It was paid for by a Wilkes Research and Scholarship Grant.
The College of Science and Engineering was recently awarded a half-million grant from the National Science Foundation to be used for a high-performance computer cluster.
Parker Dorsey is a senior communications studies major with a concentration in strategic communications. Parker began as a staff writer for the opinion...